We estimate the fraction of multiplets using 50/50 mixed cell line samples (K562/RAJI cell lines). We leverage the known genotype profiles for each of the two cell lines and calculate the cells that are assigned to ‘artificial’ genotype calls.
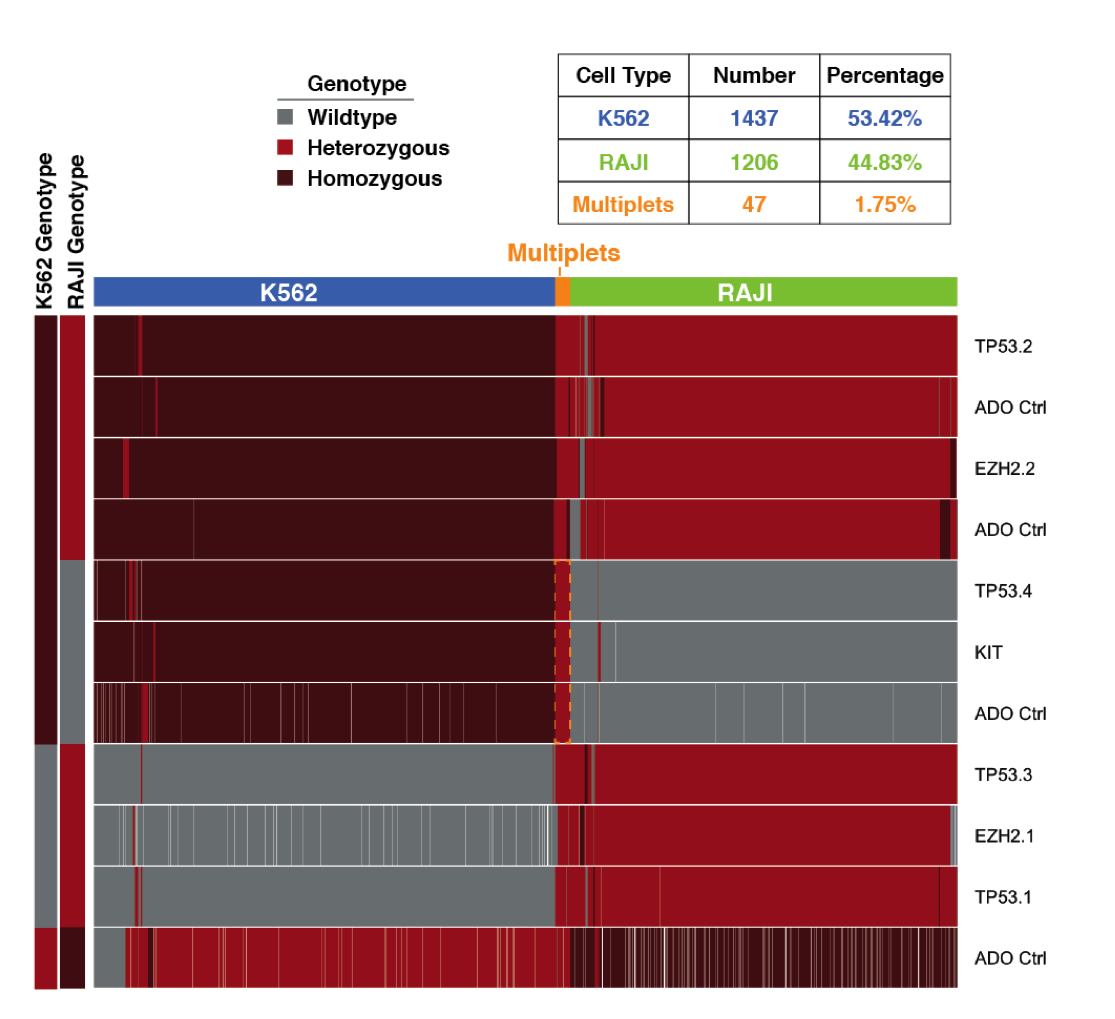
The below image shows data in a heatmap format of a representative experiment. Colors refer to genotype calls. Rows = passed-filter variants, and columns = cells.
Orange shows the fraction of cells that are inferred to be derived from droplets containing more than one cell. The 3 variants in the middle that reside on the amplicons TP53.4, KIT, ADO Ctrl are genotyped heterozygous at loci where one expects only homozygous reference (RAJI) or homozygous mutant (K562) genotype calls considering both cell lines.
The estimated doublet rate for the Tapestri Platform is < 8 %.